search this blog
Friday, March 13, 2015
Yamnaya-related ancestry proportions in Europe and west Asia
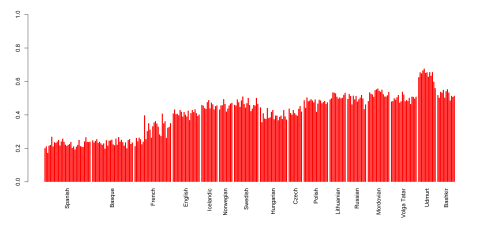
Here's a quick and dirty attempt to flush out a Yamnaya-specific ancestral component with the ADMIXTURE software and a few Yamnaya genomes from the recent Haak et al. paper: K6 spreadsheet.
Obviously, we'll need many more ancient samples from the vast Yamnaya horizon to be able to estimate direct Yamnaya ancestry in modern populations with any great confidence. But I'd say this looks like a very reasonable attempt, with more or less comparable results to those published by Haak et al. (for instance, see Figure 3 from the study here).
Please note that this wasn't a supervised run. In other words, I didn't mark the Yamnaya genomes as reference samples with the aim of creating a cluster from them.
However, I initially excluded all individuals from northeastern Europe, the north Caucasus and South Asia from the analysis. The reason I did this was because samples from these regions have a peculiar habit of creating very robust clusters in ADMIXTURE, which is useful when looking at recent variation and wanting low cross validation errors, but not so great when trying to resurrect genetic components from the depths of prehistory.
Once I had a dataset that was forcing the algorithm to focus its attention on the ancient genomes and producing consistent results, I tested the problem samples in batches of 5-10, thus making sure they didn't skew the analysis.
Interestingly, the Yamnaya-specific component peaks in Udmurts, who live close to where the Yamnaya samples were collected. This can hardly be a coincidence.
In any case, I'm hoping to look at this issue in more detail soon with the help of qpAdm, a new program released recently with the updated ADMIXTOOLS package (see here). Based on f4 statistics, qpAdm is specifically designed for analyzing ancient admixture events.
Citation...
Haak et al., Massive migration from the steppe was a source for Indo-European languages in Europe, Nature, Advance online publication, doi:10.1038/nature14317
Subscribe to:
Post Comments (Atom)

240 comments:
«Oldest ‹Older 201 – 240 of 240Where are the Italians and Greeks?
I'm guessing North Italians 25%, mainland Greeks 20% Yamna?
What do you mean?
Have you looked in the spreadsheet?
Oops, no I didn't look at the spreadsheet. Was just looking at your graphs.
Looks like I was pretty close on my estimates. Anyway, awesome analysis mate. Thanks
Matt and David,
I went and separated the German farmers by geography. I have the Stuttgart farmers SW, versus the Saxony_Anhalt NE. I didn't include the Stuttgart female from previous papers as she is older and decreased the affinity to Spain. This is all very interesting and could give credence to that Cardial/La Hoguette deal. Have a look.
German Farmers w/o “Stuttgart” in LBK_EN_SW
HungaryGamba_EN LBK_EN_SW Spain_EN Chimp -0.0144 -2.878 12494 12859 267318
HungaryGamba_EN LBK_EN_NE Spain_EN Chimp -0.0187 -3.997 12838 13326 277337
LBK_EN_SW LBK_EN_NE HungaryGamba_EN Chimp 0.0015 0.345 12592 12554 269653
LBK_EN_SW LBK_EN_NE Spain_EN Chimp -0.0061 -1.659 15599 15790 334374
LBK_EN_SW LBK_EN_NE Loschbour Chimp -0.0018 -0.406 15508 15564 334708
LBK_EN_SW LBK_EN_NE LaBrana1 Chimp -0.0001 -0.029 14717 14720 319622
LBK_EN_SW LBK_EN_NE HungaryGamba_HG Chimp -0.0026 -0.494 10775 10830 232655
LBK_EN_SW HungaryGamba_EN Loschbour Chimp 0.0092 1.525 12679 12448 267468
Spain_EN HungaryGamba_EN Loschbour Chimp 0.0224 3.968 13119 12544 275071
HungaryGamba_EN LBK_EN_SW Spain_EN Chimp -0.0144 -2.878 12494 12859 267318
HungaryGamba_EN LBK_EN_NE Spain_EN Chimp -0.0187 -3.997 12838 13326 277337
LBK_EN_SW Iceman LBK_EN_NE Chimp 0.0030 0.565 15530 15438 335725
LBK_EN_SW Iceman HungaryGamba_EN Chimp -0.0088 -1.340 12128 12344 268408
LBK_EN_NE Iceman HungaryGamba_EN Chimp -0.0100 -1.694 12546 12798 279586
LBK_EN_SW Iceman Spain_EN Chimp -0.0066 -1.243 15133 15336 332654
LBK_EN_NE Iceman Spain_EN Chimp 0.0006 0.124 15788 15768 345684
Keep in mind that the SW samples are older than the NE samples by a couple hundred years. Also, removing "Stuttgart" increased the Z score by -.4, so it was fairly important to showing the change.
@Mike and Maju
Thanks for your comments.
Is there any better information about Cernavoda than Mallory's short comment about it? If the dating there is correct (4000-3200 BCE) then it mostly precedes Yamnaya, yet it's a Kurganized culture (and even when it overlaps with Yamnaya, the CT culture was in between both, so direct contact is not granted).
It's strange that it hasn't been studied or dated better, being the first Kurgan culture in Europe (as far as I can tell).
@Chad
Thanks for all those stats. It seems you're trying to squeeze out all the smallest details out of those genomes.
In the Haak paper I found something intriguing. The Fst table on page 26 shows that all modern European pops are closer to EHG than to WHG, including Basque and Spanish. This seems a bit odd.
Would you be able to try something like:
Loschbour Samara_HG Test Chimp
Where Test are European populations (Spanish, Basque, French, German, Polish, Lithuanian, Tuscan, Greek...)?
Thanks!
@ Alberto
Excatly. Cernadova dates from 4000 BC. So it cannot be a "Kurgan" culture unless we start bending the picture. Rather, it is as I ave long been stating - a late phase local variant of the Cucuteni culture. If one examines closely traditional archaeology, the Cucuteni disperals imparted a significant colonization affect onto the western steppe. Genetic studies will prove/ disprove/ modify this picutre.
See Eg : IGOR MANZURA
STEPS TO THE STEPPE: OR, HOW THE NORTH PONTIC REGION WAS COLONISED (Igor Mazura).
There is a book "IMPORT AND IMITATION
IN ARCHAEOLOGY" witha few good chapters.
Theres an 'old skool' perspective by Hoddinott. "From the Chalcolithic to the EBA in west and north Pontic lands"
There are numerous papers on the Baden Cotofeni horizon.
In fact, just give me your email; I have a whole dropbox file worth of papers..
THE PROTO-BRONZEAGE CEMETERYAT DURANKULAK: A LOOK FROM THE EAST
Focuses on Cernadova
Yes, CT must have had a good influence in the western steppe.
Thanks for all the references. I think it's enough for now to get me started. If I need something else I'll ask, thanks :)
Almost Forgot
Phil Kohl:
"Crossing the Pastoral/ Agricultural bridge"
Alberto,
I'll give those a crack this afternoon, after work.
@Chad: OK, I seem to have misinterpreted the abstract.
@David "I'd say some ANE also entered Europe from west Asia during the Iron Age and middle ages, and was very likely accompanied by a very low ratio of EHG and a high ratio of Near Eastern ancestry".
Not sure of which parts of Europe do you mean nor why this ME+ANE could not have entered Europe (initially only the Balcans) c. 5000 BCE, which is the Vinca arrival time-frame. Otherwise I only see windows for Italy in the Bronze Age mostly and maybe something Phoenician (???) in the case of Sicily and Malta. But nothing affecting Europe in general, only the SE.
Another thing would be if some of that Italian complex would have scattered around in Roman Imperial context (?), but still not likely to affect Northern nor Eastern Europe for example. Also I don't see any obvious data supporting any major Roman-related genetic expansion.
@Krefter: the North African element in West Iberia is well known and has no relationship whatsoever with the extra West Asian element we are discussing here. As for East Iberian much smaller extra ME, IMO fits better with Italian colonial overflow but nothing big in any case.
All this could well be related to Y-DNA J2 and IMO this could well have arrived to SE Europe as early as 5000 BCE with Vinca-Dimini (regardless of possible secondary flows such as Etruscans).
Interesting point by Jean continued by Maju - the Basques and Spanish do deviate the general relationship between Yamnaya-related and ANE K8:
e.g. graphed against one another for the West Eurasian samples which overlap http://i.imgur.com/IxmGpXy.jpg
Two clines... The Middle East and West Asian has an aversion to the Yamnaya component.
With the Pathans and Turkmens included, the Pathans are just above the Lezgins (more ANE relative to Yamnaya than Lezgins) and Turkmens just below (less ANE relative to Yamnaya than Lezgins). But that's less unexpected than Southwest Europeans slightly bucking the relationship between ANE and Yamnaya.
@Alberto, the WHG are further from Europeans because of their high levels of drift overall -
e.g. WHG-Yoruba FST is 0.195, while EHG-Yoruba is only 0.179. WHG is just more drifted here.
Similarly, Baalberge_MN-EHG is 0.067 while Baalberge_MN-WHG is 0.073, despite the fact Baalberge is clearly more related to WHG by f4.
However, the stats you propose for Europeans would be a good supplement to the p73 stats assessing the same relationship in ancient Europeans.
@Matt: very informative graph, thank you.
I can't but notice that your clinal line, while apparent, does not express a normalized ratio of ANE:Yamna, because this would have to drop to 0:0. I drew a few reasonable examples of such ratios in blue color in this annotated version of your graph.
the Volga-Ural pop. seems to be in the 0.04 (ANE for each 1 Yamna score, i.e. 1:0.04) ratio, while the Basque-Iberian one is in the 0.03 instead, however other populations have much larger ratios: 0.09 and 0.18 drawn for reference but even 1:0 in some Near East (?) cases, which have no Yamna but significantly strong ANE. On the opposite extreme are Sardinians who vary on the Yamna side but always score zero for ANE (0:1 ratio).
So I gather that, unless I'm reading something wrong, ANE and Yamna only have a weak correlation. Possibly various ancestral populations carried different scores of Yamna-like and ANE-like genetics, causing all this mess. Very apparently the ones influencing West Asia had much more ANE than Yamna, while the ones influencing SW Europe had much more Yamna than ANE.
The first one is easy to explain: ANE affinity in West Asia has nothing (or just a little) to do with IEs but the component was there all the time, at least in bulk. However the latter is harder to explain: either Lochsbour/La Braña or maybe Gökhem fit in there (extra HG unaccounted for) or else we have to appeal to some ghost population.
Erratum: "ANE for each 1 Yamna score, i.e. 1:0.04" should read "ANE for each 0.1 Yamna score, i.e. 0.1:0.04" - or maybe I should have just said: "10:4".
Anyhow is it possible that the scores can't be considered absolute and that there's a possible "real {0,0}" to the left of the graph, in the "negative Yamna" zone, approx at the -0.3 position? The tendencies suggest so and, if so, the Yamna:ANE mystery could be conciliated somehow.
In the event that this "displaced {0:0}" theory is correct, I drew this other version of the graph, which suggests four cases:
1. The Reference case, which follows the main cline and is similar, although not exactly so, to your red line.
2. Caucasus-Zagros: clearly more ANE than Yamna rel. to the Reference ratio.
3. Iberia (incl. Basques): clearly more Yamna than ANE rel. to the Reference ratio.
4. Sardinia: all Yamna and no or nearly no ANE whatsoever.
So this is the question I have asked often when people say "X pop. has N ANE" or stuff like that: compared to what? It's an affinity degree not an absolute thing!
I got caught up tonight. I will try the ancient farmers and such, with f4 tomorrow. Sounds interesting.
@Matt
Yes, that drift you mention might be the cause, but ?d just thought that the drift was shared with the populations that descend from them. But probably f4 stats will show the expected results instead of the unexpected Fts values.
As for that Yamnaya/ANE ratios, wouldn't the explanation be related to WHG/ANE ratio? Spanish and Basques are the one who have a higher WHG/ANE ratio while Caucasus/NE/Central-South Asians have the lowest. In the rest of Europe there is a good correlation between both: population with highr WHG also have higher ANE (Baltics), while population with low WHG (SE Euros) have lower ANE.
@Chad
Thanks, I'd appreciate if you give those a try.
Matt and Alberto,
result: Karelia_HG Loschbour LaBrana1 Chimp -0.1296 -16.786 13350 17325 320734
result: Karelia_HG Loschbour LaBrana1 Ust_Ishim -0.1279 -14.781 12581 16270 320085
result: Karelia_HG Loschbour HungaryGamba_HG Chimp -0.1181 -13.020 9930 12590 233238
result: Karelia_HG Loschbour HungaryGamba_HG Ust_Ishim -0.1161 -12.252 9344 11800 232756
result: Karelia_HG Loschbour Clovis Chimp 0.0547 6.765 16436 14730 337725
result: Karelia_HG Loschbour Clovis Ust_Ishim 0.0671 7.621 15306 13382 337042
result: Karelia_HG Loschbour Ust_Ishim Chimp -0.0070 -0.936 15140 15354 337801
result: Karelia_HG Loschbour Kostenki14 Ust_Ishim -0.0202 -2.290 13353 13902 317861
result: Karelia_HG Loschbour HungaryGamba_EN Chimp -0.0267 -3.594 12175 12843 270325
result: Karelia_HG Loschbour HungaryGamba_EN Ust_Ishim -0.0228 -2.779 11520 12057 269786
result: Karelia_HG Loschbour LBK_EN_SW Chimp -0.0188 -2.785 15123 15703 327098
result: Karelia_HG Loschbour LBK_EN_SW Ust_Ishim -0.0116 -1.597 14198 14532 326447
result: Karelia_HG Loschbour LBK_EN_NE Chimp -0.0179 -2.870 15588 16155 338296
result: Karelia_HG Loschbour LBK_EN_NE Ust_Ishim -0.0121 -1.821 14581 14937 337612
result: Karelia_HG Loschbour Spain_EN Chimp -0.0275 -4.460 15293 16158 335494
result: Karelia_HG Loschbour Spain_EN Ust_Ishim -0.0220 -3.189 14402 15050 334814
result: Karelia_HG Loschbour Bell_Beaker_LN Chimp -0.0132 -2.181 15745 16166 335723
result: Karelia_HG Loschbour Bell_Beaker_LN Ust_Ishim -0.0069 -0.994 14829 15034 335047
result: Karelia_HG Loschbour Corded_Ware_LN Chimp 0.0185 2.998 16405 15809 338141
result: Karelia_HG Loschbour Corded_Ware_LN Ust_Ishim 0.0270 3.753 15341 14536 337458
result: Karelia_HG Loschbour Karsdorf_LN Chimp 0.0084 0.607 2808 2760 58388
result: Karelia_HG Loschbour Karsdorf_LN Ust_Ishim 0.0251 1.842 2646 2516 58262
result: Karelia_HG Loschbour Motala_HG Chimp -0.0297 -4.701 15618 16574 334629
result: Karelia_HG Loschbour Motala_HG Ust_Ishim -0.0242 -3.318 14776 15510 333952
result: Karelia_HG Loschbour SwedenSkoglund_NHG Chimp -0.0358 -4.495 14450 15522 311225
result: Karelia_HG Loschbour SwedenSkoglund_NHG Ust_Ishim -0.0305 -3.672 13728 14591 310596
result: Karelia_HG Loschbour Samara_HG Chimp 0.0622 7.260 10251 9051 201624
result: Karelia_HG Loschbour Samara_HG Ust_Ishim 0.0735 7.883 9695 8366 201206
Thanks you!
The ones that better relates to what I was wondering are:
Karelia_HG Loschbour Bell_Beaker_LN Chimp -0.0132 -2.181
Karelia_HG Loschbour Corded_Ware_LN Chimp 0.0185 2.998
So Basically CW is closer to EHG than to WHG, but BB is closer to WHG than to EHG.
So I guess it's similar with modern populations. The cut might be around Poland: east and north of Poland they might be closer to EHG, but south and west closer to WHG.
So, Chad and Alberto: do those tests say or at least suggest that part of the Yamna component is actually Lochsbour-like (WHG)? And, for what I understand, also that Bell Beaker people of Germany had it rather than actual Yamna admixture?
I wouldn't read it that way.
I mean, Yamnaya does have WHG (some 35%), but it comes mostly from the EHG (who had more than 50% of it).
So basically Europeans have WHG from 2 sources: directly from Loschbour-like people, and from Yamnaya.
So places with very high Yamnaya admixture are closer to EHG than to Loschbour, while as Yamnaya admixture decreases, it turns the other way around.
Basically the results are as expected, and what was wrong was that table in the paper showing ALL Europeans closer to EHG than to WHG (including for example, Basques, which was very odd). But Matt explained above the reason that that oddity and Chad's test confirm that it was indeed a false clue.
But then, Alberto, how do you explain that the Yamna-ANE correlation is so varied: that some peoples (particularly SW Europeans) have clearly more Yamna fraction that should correspond to their ANE fraction? Both Yamna and EHG are high in ANE yet when we found those same components in the west (per this run by Davidski or also per Haak et al.) it seems to have "lost" a good deal of its ANE (all in the case of Sardinians)?
I would rather postulate a "hidden" HG component that is not being evaluated. A component that would be Lochsbour-like in ANE but otherwise Yamna-like (or EHG-like) enough to pass as such.
Yes, now I see your point.
I thought it was a simple question of ANE/WHG ratio, but it's not so simple. As you point out, Yamnaya seems to be picking signal from extra WHG in relation to the amount of ANE.
I really have no idea why it happens.
Another possibility is that, judging on the K=16 of Haak et al., Yamna is expressed as a 50-50 mix of HG and what we can call "West Asian 2" (WA2, dark grayish green), which is in essence: all West Asian minus the EEF component (orange).
In general terms both WA2 and HG (plain blue) have more ANE than the EEF component but this relation may vary and we can't measure it in general terms because Ma1 has no effective influence on the analysis (too old and too small sample to make sense in such a large Holocene sample). However Yamna is always expressed as 50-50 HG+WA2, picking signals from both these "more true" components regardless of ANE affinity and true Yamna ancestry. In essence: every time WA2+HG is found, it's classed as Yamna in Davidski's run, regardless of the suptypes of HG and WA2 involved and regardless of ANE affinity (with the remaining HG cast as "other HG").
Hence we can surely use the differential ratios to estimate various scenarios, depending on how exactly Yamna performs analyzed against itself and ANE. Throwing there also other ancient pops like CW or BB could be interesting.
Matt: do you think you can do that?
Sergey,
Is there any chance you can also get decent files of Starcevo_EN and LBKT_EN?
Combining them might make a very useful early Neolithic reference for populations not in the Human Origins dataset. They're about as close as we've got now to Neolithic farmers from the Near East.
Unfortunately, no. BAM files for LBKT and Starcevo are tiny. All appropriate files have been processed now.
Just an observation from this study that M269 freq., Lezgins is the highest and is the one closest to Yamnaya as per Shaikorth. What about Ossets–Digor?
R1b1b2-M269
(supplementary fig. 1, Supplementary Material online) were
found in the Lezghins (30%) and in Ossets–Digor (16%).
Parallel Evolution of Genes and Languages in the Caucasus Region
Balanovsky et al
August 27, 2011
http://mbe.oxfordjournals.org/content/28/10/2905.abstract
Alberto said...
"Good idea, although Lezgins have considerable overlap with Yamnaya because of their particular mixture."<Shaikorth said...
...
For the results, Tuscans don't seem to like Yamnaya much, but Lezgins are not the best fit either (as source of ANE).
March 14, 2015 at 6:09 AM
Davidski said...
"Gill,
Yes, I think so, otherwise we'd see much higher Yamnaya among South Central Asians, inflated by the high ANE among them, but we don't. There's a nice gradient there from the Volga-Ural to the Pamirs and into the Hindu Kush and Iran. It just makes sense.
...
March 15, 2015 at 4:28 PM
MJost
Alberto: As for that Yamnaya/ANE ratios, wouldn't the explanation be related to WHG/ANE ratio? Spanish and Basques are the one who have a higher WHG/ANE ratio while Caucasus/NE/Central-South Asians have the lowest. In the rest of Europe there is a good correlation between both: population with highr WHG also have higher ANE (Baltics), while population with low WHG (SE Euros) have lower ANE.
Alberto, that's a good point and it could do.
Although I would expect this should have fallen into having additional WHG_extra cluster relative to their K8 ANE and not Yamnaya, if the test is actually picking up WHG as it should. That's supposed to be what allowing WHG to vary achieves.
Maju: I can't but notice that your clinal line, while apparent, does not express a normalized ratio of ANE:Yamna, because this would have to drop to 0:0. I drew a few reasonable examples of such ratios in blue color in this annotated version of your graph.
Good point.
Tentatively, part of the way both your comments seem to resolves, and the way that the Basque and Spanish have Yamnaya than expected from ANE K8, is that probably some level of what is covered by the ANE cluster in K8 is in the Middle Eastern cluster, here.
When I mentioned Levantines on the other, graph, I really overstated matters - note that there are few Levantines in the K6 run, and the ones that are (Lebanese Christians, Syrians, Jordanians) have lower membership in the Middle Eastern cluster than the Armenians, Georgians and Iraqi Jewish.
Basques lack membership in the Middle Eastern cluster.
Some visual circumstantial evidence is, if you PCA only the Yamnaya K6 measures of Yamnaya_related and Middle_Eastern, and then PCA them together with the ANE measure, you can see where they point.
http://i.imgur.com/UBHUrpb.jpg - typical West Eurasian groups only
http://i.imgur.com/saSrT1U.png - plus Pathans, Turkmens, Tatars, Kazakhs
The Sardinians, Basques, Spanish, Caucasus, Near East, Levant, North Europeans, etc are all where you expect them to be. Middle Eastern points to West Asia.
So seems, the Middle Eastern cluster here is not a measure of ANE free Middle Eastern ancestry, however we interpret what it does measure and whether it is real cluster of similar age to pre-Yamnaya and Yamnaya, and how we interpret the idea of ANE.
Btw, a measure of K6_ANE_estimate =(Yamnaya_Related*0.32)+(Middle_Eastern*0.17) seems to give the best correlation with K8 ANE at 0.94 across the samples in common, and the average difference between them per sample is *pretty* close to 0%, median difference 1% (Basques are still around 2% wrong but its the best fit I can find).
Slight differences are still Sardinians, early Neolithic and Pathans / Tajiks at the edges (S are over by 5%, P & T are under by around 10%), but it's almost exactly the same for most samples.
http://i.imgur.com/YD7i5ln.png
The Middle_Eastern cluster here seems pretty much close to Georgians, but not quite.
@Matt: that's a very impressive fit, thank you for the work, really.
It would seem, judging on your work, that we are before two (main) different sources of ANE affinity: Yamna can account for ~2/3 but there is another source that is best described as Highland West Asian, accounting for ~1/3.
Ironically the best fit for Yamna only influence are Basques. That has me very perplex really, more so when mtDNA wise the Basque genetic pool seems very stable since Neolithic times (Kurgan-derived influences here before the Iron Age are extremely unlikely and most likely negligible beyond the normal neighbor exchange, what should be very diluted). I still think that there's something fishy re. what appears as Yamna genetic influence.
PS- well, the best fit for Yamna-only influence in ANE would be Sardinians, admittedly. Still fishy.
Not if you get out of R1b megalith mind.
Maju, I don't know much about how the Basque genetic uniparental pool has changed over time (as we move from archaeology from people living where the Basques live to people who we can identify clearly as Basuqe) still I think those sound like pretty solid points.
Btw, for similar formula for WHG and Near Eastern, the closest I could get were:
K6 WHG estimate = WHG_Extra+0.38*Yamnaya_Related+0.41*Pre_Yamnaya
K6 Near_Eastern estimate = 0.3*Yamnaya_Related+0.59*Pre_Yamnaya+0.83*K6_Middle_Eastern
These have 0.99 and 0.98 correlations with the K8 components, and the mean and median differences between each sample on them are low.
Although, even though the correlations work out and the average differences are low, the populations end up less different from one another on the estimated components than in K8.
It seems like at least some populations who are extreme relative to others can have differences from one another in calculators with different / higher K compared to K8, e.g.
Neolithic Hungary and Stuttgart in Yamnaya_K6 here has some of the Middle_Eastern component so fit with the formula should have ANE but doesn't in K8 (I think)
Sardinians in Eurogenes K15 have some North_Sea so would be predicted to have some ANE based on North_Sea having ANE but don't in K8
Lithuanians would have more Near_Eastern based on the above estimate than in K8, etc.
Even the K15 Atlantic component has noticeable ANE according to TreeMix tests, and that is common in modern North Europeans but also in Sardinians and even Early/Middle Neolithics.
However, in other calculators like K36 the neolithic farmers distinctly lack the Atlantic coast-associated components (North Atlantic and North Sea) that are especially present in Northwest Europeans who have lots of K15 Atlantic.
The lesson I'd take from this is to rely less on ADMIXTURE results and more on formal testing, even PCA positioning.
@ Davidski
Could you please add Belarusians and Ukrainians to the spreadsheet?
@Davidski
Can you please add South Asian samples you have to this spreadsheet?
Are these calculations still reliable? (After new datas and methods)
Post a Comment